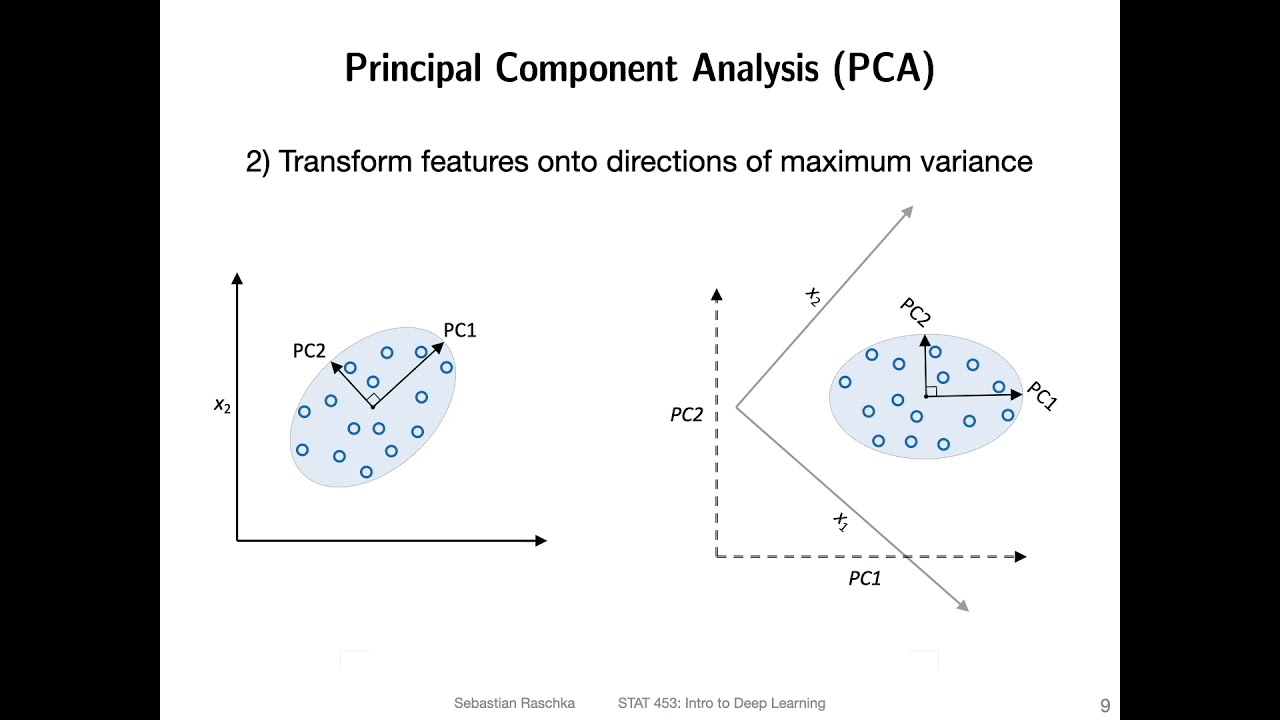
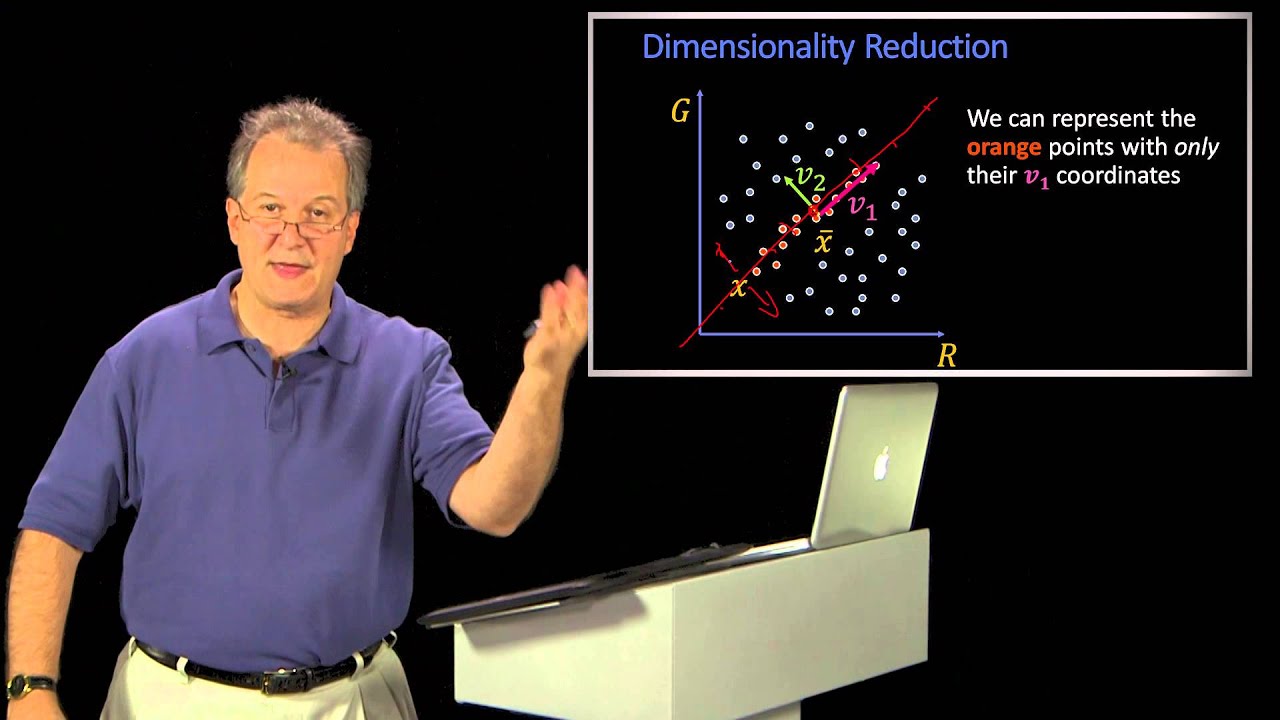
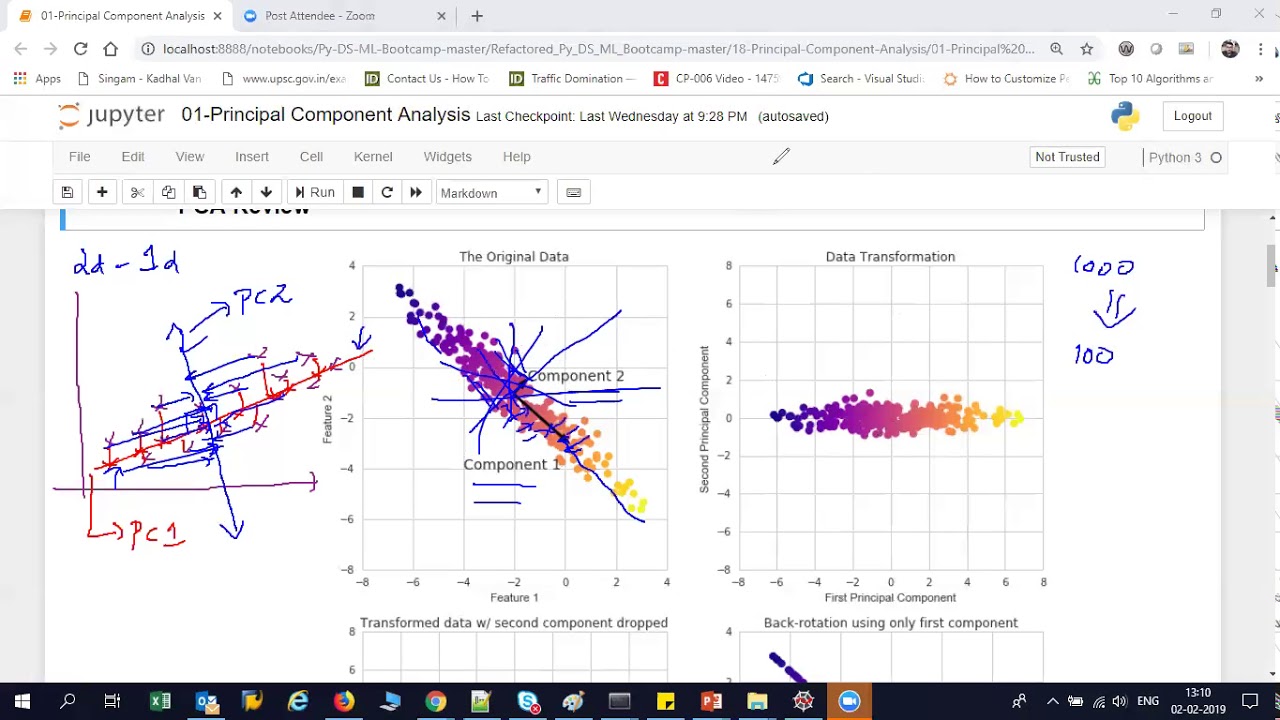
foreign which is dimension reduction and so if we remember in the subset selection methods we were just taking a subset of the predictors and using least squares to fit the model and then in Ridge regression on the lasso we were we were really doing something different where we were taking all of the predictors but we weren't using least squares we were using a shrinkage approach to fit the model and now we're going to do something different which is we're going to use least squares but we're not going to use least squares on the original predictors
X1 through XP instead we're going to come up with new predictors which are linear combinations of the original predictors and we're going to use these new predictors to fit a linear model using least squares so this is known as Dimension reduction and the reason it's called Dimension reduction is because we're going to use those P original predictors to fit a model using M new predictors where m is going to be less than P so we're going to shrink the problem from one of P predictors to one of M predictors so in a little bit of
detail here we're going to Define m linear combinations Z1 through z m where m is some number less than p and these are going to be linear combinations of the original p predictors so for instance ZM is going to be the sum of the P predictors where each predictor is multiplied by Phi MJ where vmj is some constant and in a minute we'll talk about where this vmj comes from but the point is once we get our new predictor is Z1 through z m we're just going to fit a linear regression model using least squares
but instead of using the X's we're going to use the Z's so in this newly squares model my predictors are going to be disease and my coefficients are going to be Theta naught through Theta m and the idea is that if I can just be really clever in how I choose these linear combinations in particular if I'm clever about how I choose these vmjs then I can actually beat least squares that I would have gotten if I had just used the Raw predictors so um one thing that we should notice is that you know on
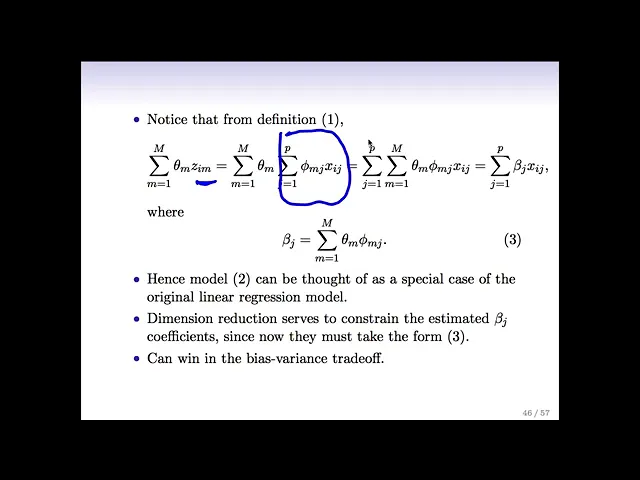
the previous slide here we had this this summation over Theta m z i m and if we look at that a little bit more carefully and we plug in the definition of Zim which remember was just a linear combination of the original X's and we we switched to the order of the sums and we do a little bit of algebra we see that what we actually have here is a sum over the p predictors times this quantity times the pth predictor so this is actually just a linear combination of the original X's where the linear
combination involves a beta J that's defined like this so the point is that when I do this Dimension reduction approach and I Define these new Z's that are linear combinations of the X's I'm actually going to fit a linear model that's linear in the original X's but the linear the the beta JS in my model need to take a very very specific form so so these Dimension reduction approaches they're giving me models fit by least squares but but I'm fitting the model not on the original predictors it's on a new set of predictors and I
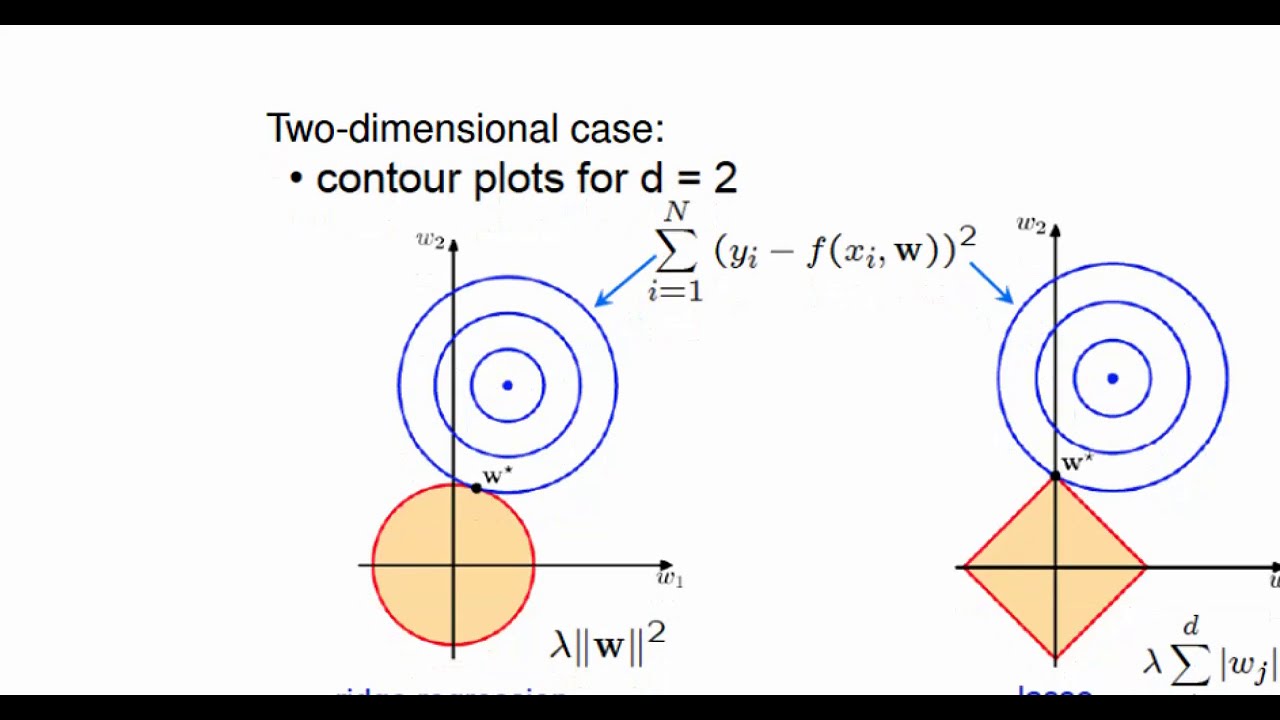
can think of it actually as ultimately a linear model on the original predictors but using different coefficients that kind of take this funny form here so in a way it's sort of similar to Ridge and lasso right it's still least squares still a linear model in all the variables but there's a There's a constraint on the coefficients that's exactly right but we're getting a constraint in a different way we're not getting a constraint like in the ridge Case by saying okay my sum of squared beta is needs to be small instead we're saying my betas
need to take this really funny form if you look at it but it's got a simple interpretation in terms of least squares on a new set of features so and the the idea here is really it boils down to the bias variance trade-off by saying that my betas need to take this particular form I can win um I can get a model with low bias and also low variance relative to what I would have gotten if I had just done plain vanilla Lee squares on the original features um one thing that I should mention is
that this is only going to work nicely if m is less than p and instead if my M equaled p then I would just end up with least squares and this whole dimension reduction thing would have just given me least squares on the raw data