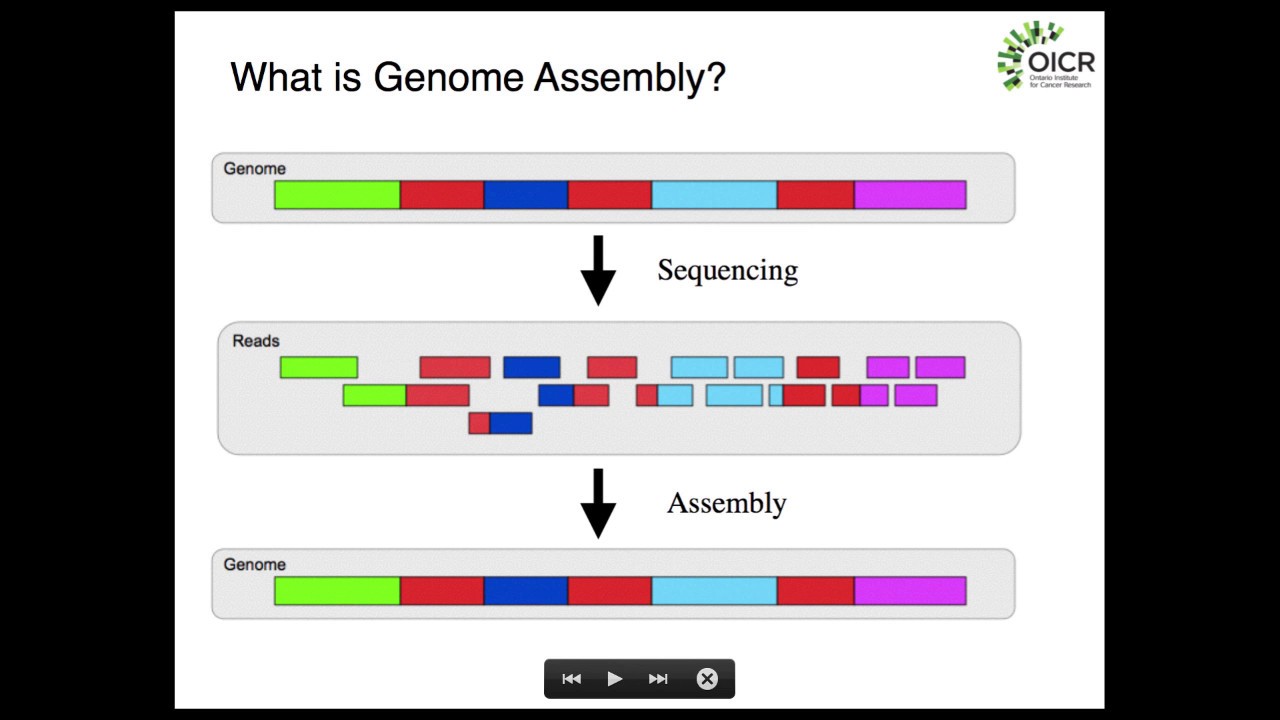
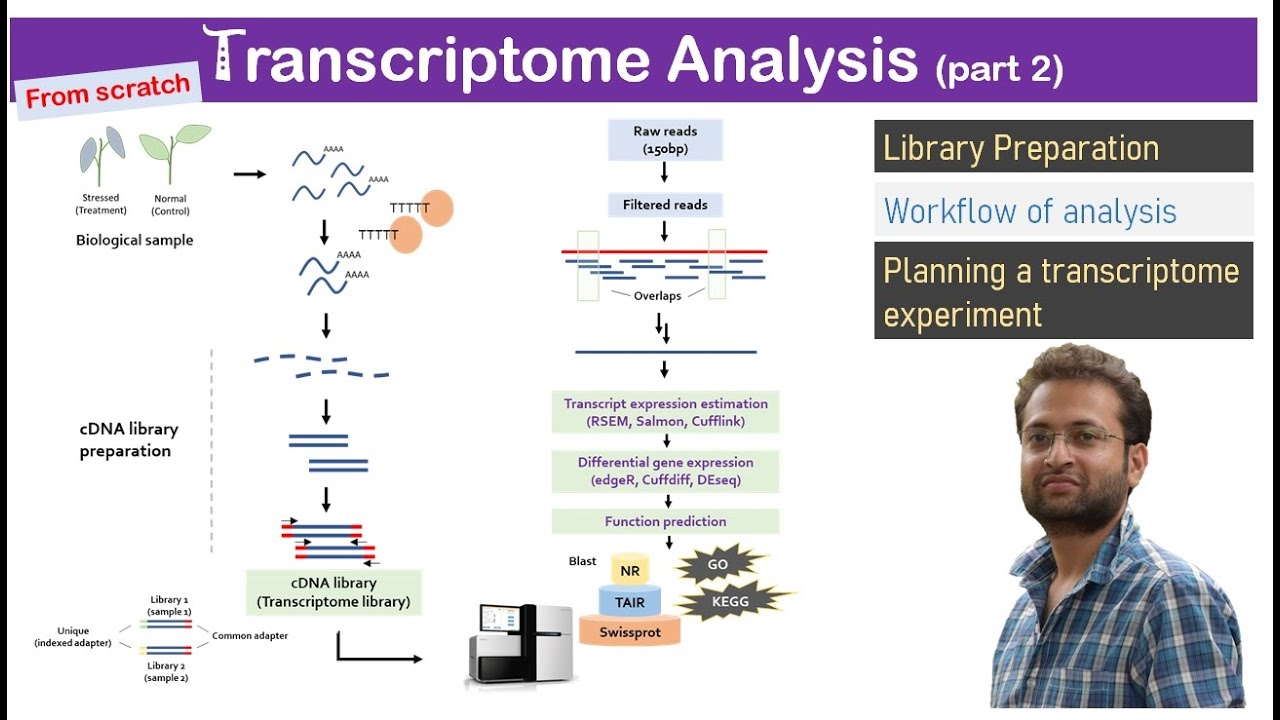
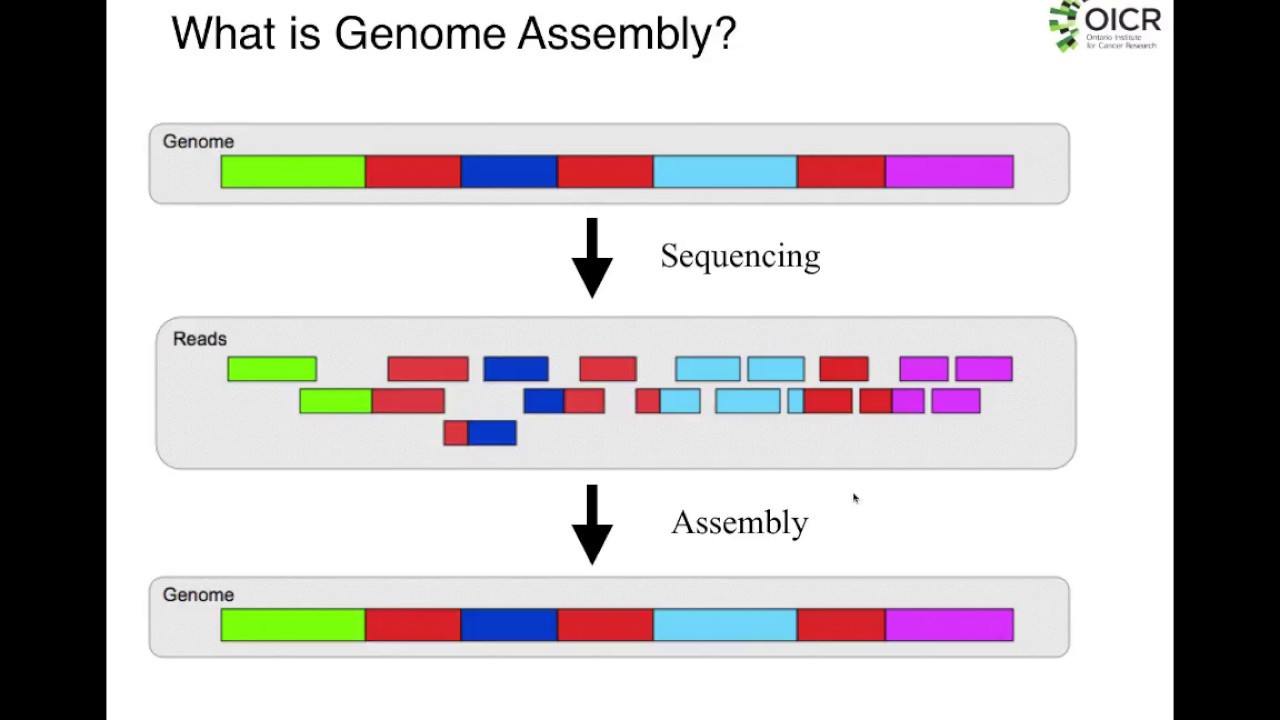
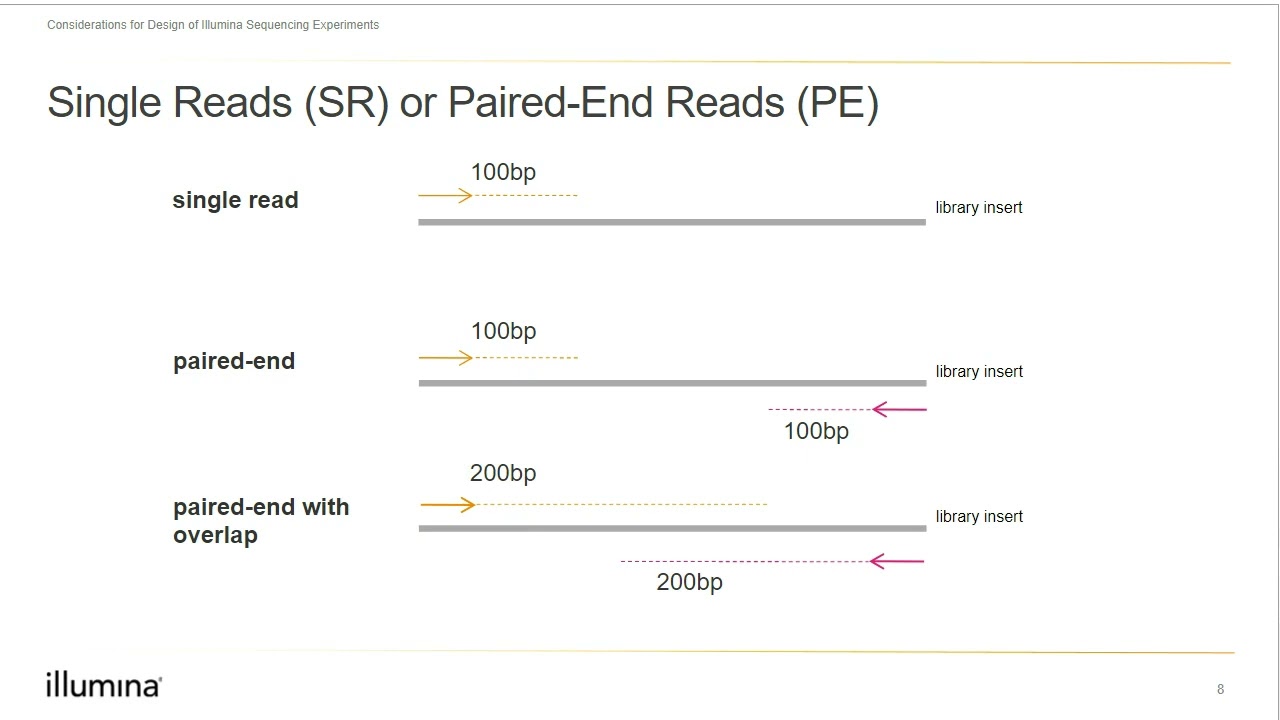
hello and welcome back to buy informatics series of explore bio and in today's video you will learn in very simple terms what a dinovo assembly is what are the major challenges in Dino assembly and some of the most popular dinovo assemblers used for assembling the genome and transcriptome genome and transcriptome of most eukaryotic organisms are huge and complex to be entirely sequenced in a single go in order to sequence the entire genome or transcriptome most sequencing approaches follow shotgun sequencing procedure and this the large fragments of DNA or complementary DNA is first broken down to
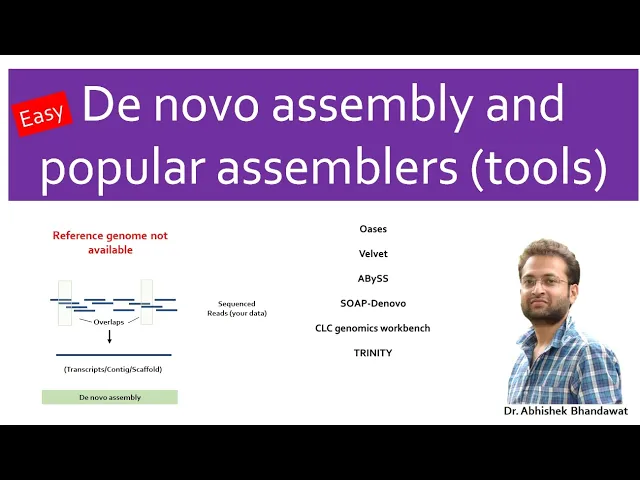
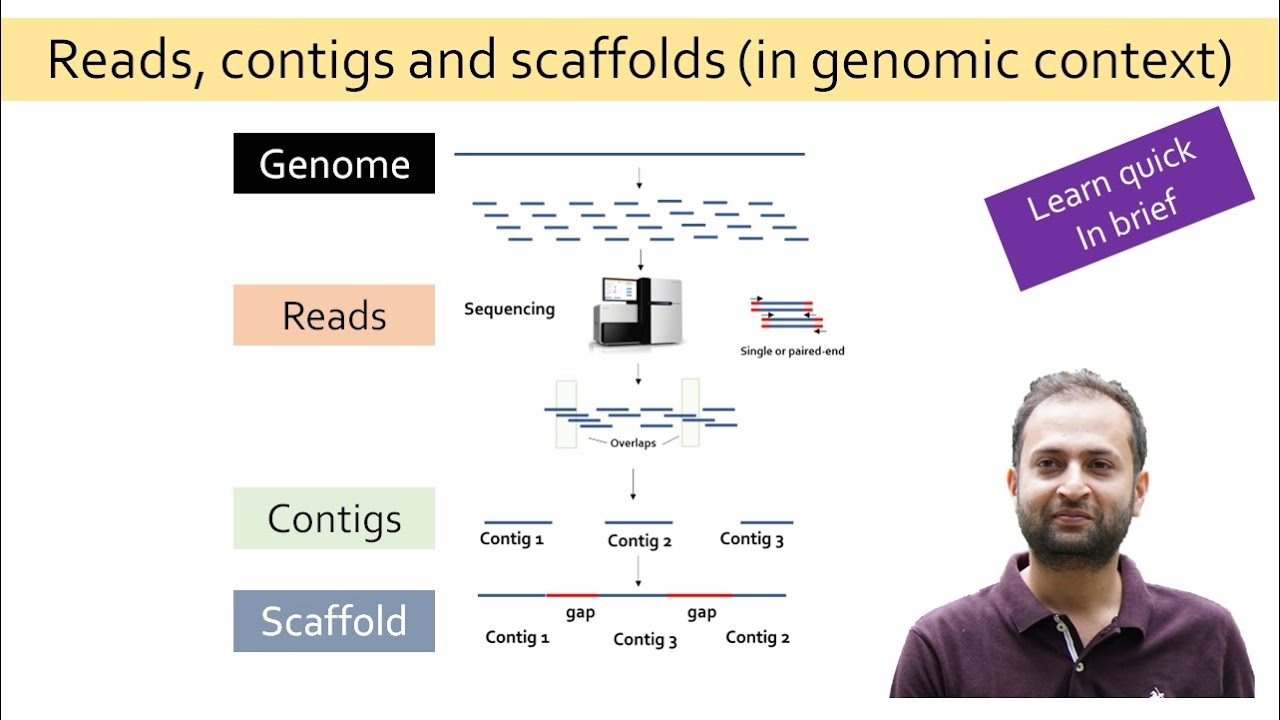
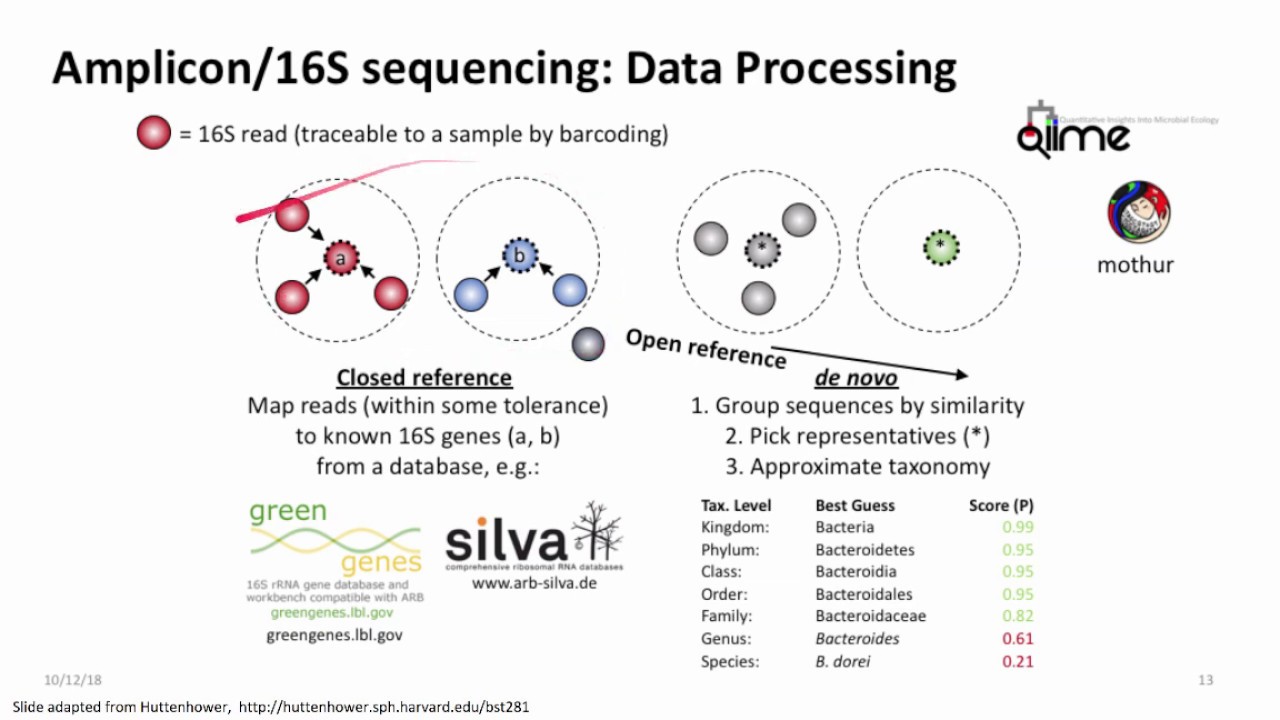
short chunks which can be sequenced by a sequencing instrument after the sequencing is completed these short fragments which are called as reads are aligned and assembled based on overlapping regions to form larger fragments which is the reflection of entire genome or transcriptome when you already have a reference genome sequence of an organism available you can map the sequence reads generated in your experiment to that already known reference sequence this is called as reference based assembly but what if you do not have a reference genome sequence for your organism of Interest available you still can align
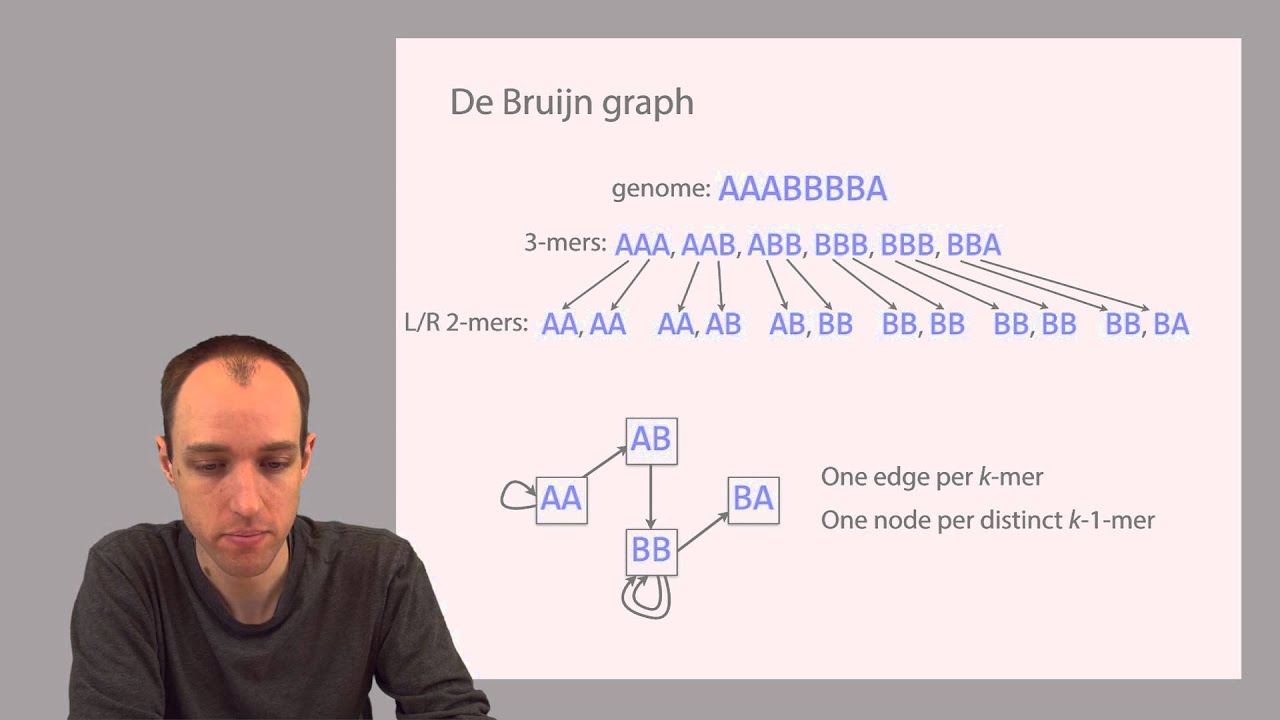
and assemble the sequences based on the overlapping regions of the reads generated without a reference thus assembling the reads to longer contigues is scaffolds and chromosomes without the reference information is termed as Lenovo assembly the meaning of denovo is from the very beginning such sequencing is termed as denovo sequencing the sequence assemblers utilizes different set of algorithms to precisely assemble the reads to transcripts or contacts or scaffold or chromosome based on minimum number of overlapping bases at the end of each raid also known as Kmarts distance between the reads in paired and reads and incorporates
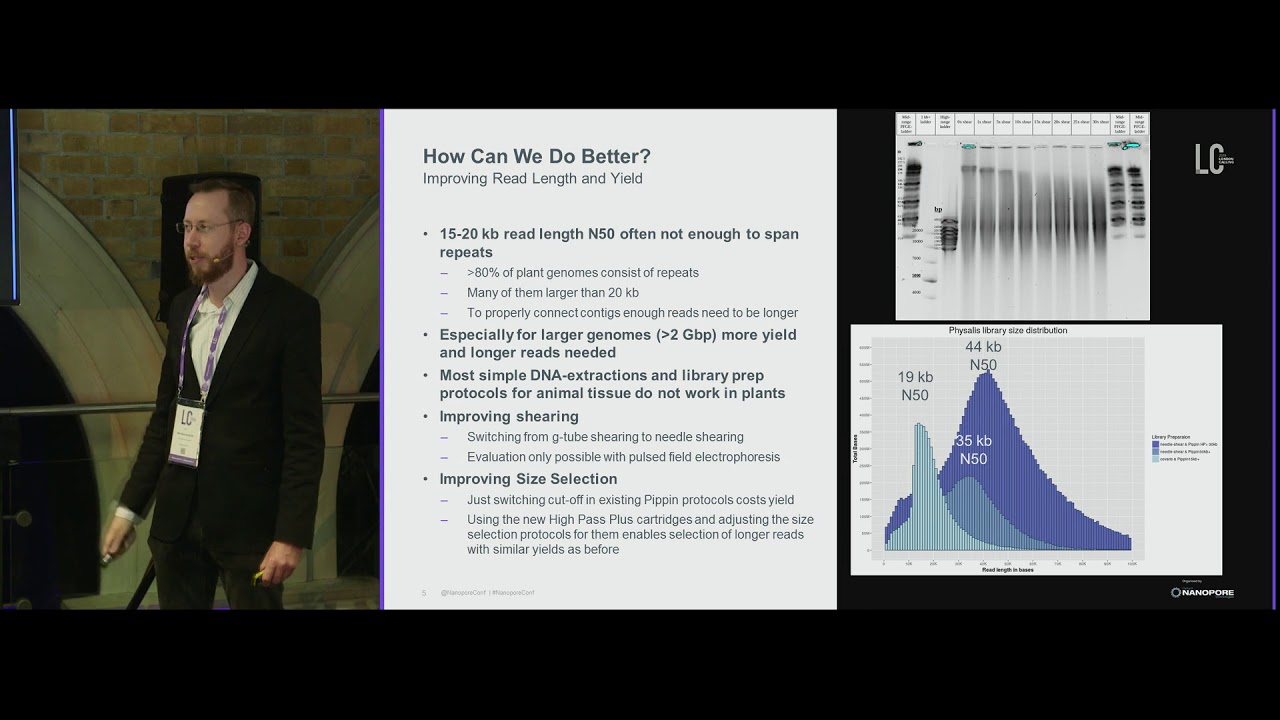
gaps if required filling in these gaps may require longer read sequencers such as spec bio and nanopore for large genome assemblies a combination of short and long read sequencing is often seen more useful coming on to the major challenges faced during the denovo assembly huge complex polyploid genome with lots of repetitive DNA and heterozycosities is the major challenge in the accurate sequence assembly errors in sequencing low genome coverage and low depth of sequencing are some of the other problems that needs to be considered while going for denovo sequencing OSS velvet app is so denovo CLC
genomics workbench Trinity are some of the most popular sequence assembly tools each of these have their own advantages and limitations that needs to be considered before selecting one for your study if you find the video useful do share it with others do check out my playlist on bioinformatics Research publishing markers and others comment below for your queries and requests I usually respond to them subscribe and hit the Bell icon to stay informed about my latest uploads [Music]