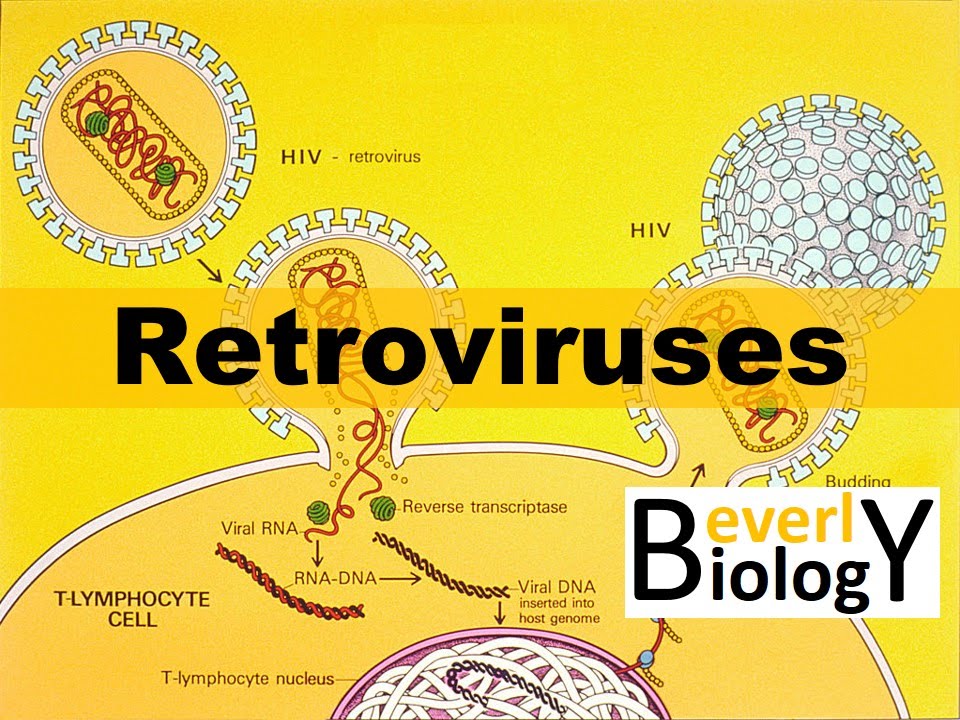
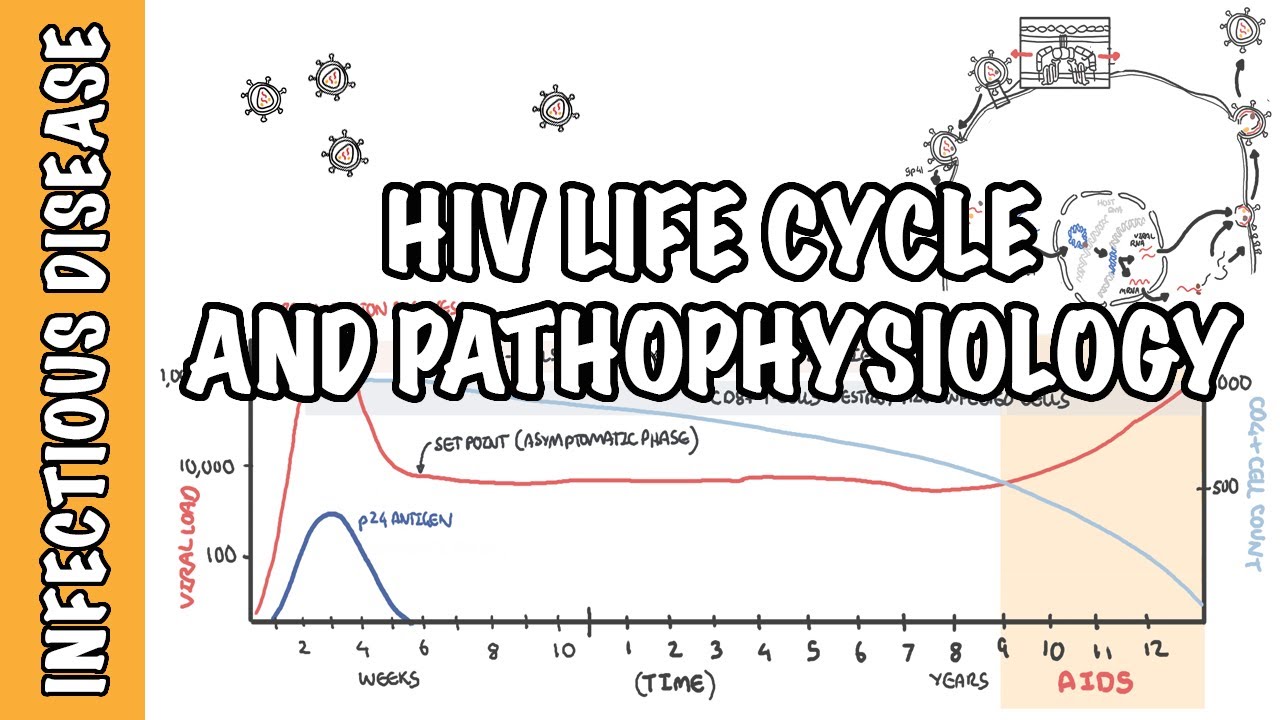
Hello, I'm Dr David Spach from the University of Washington. In this National HIV Curriculum Mini-Lecture, I'll provide an overview of the HIV life cycle. As shown in this overview diagram, HIV has a fascinating and highly complex replication cycle.
In this talk, I'm going to explore each of the major steps of the HIV life cycle as shown here. Gaining clear insights into each of these steps in the HIV lifecycle will provide the foundation for understanding the mechanism of action of the antiretroviral medications. Now, let's take an in-depth look at the first step in the HIV lifecycle, HIV entry.
This initial step involves a complex interaction between the HIV envelope glycoproteins and the host cell surface receptors. In this close-up view, HIV is shown on the top and the host cell is shown on the bottom. This illustration shows HIV approaching the surface of the cell, essentially like a lunar landing.
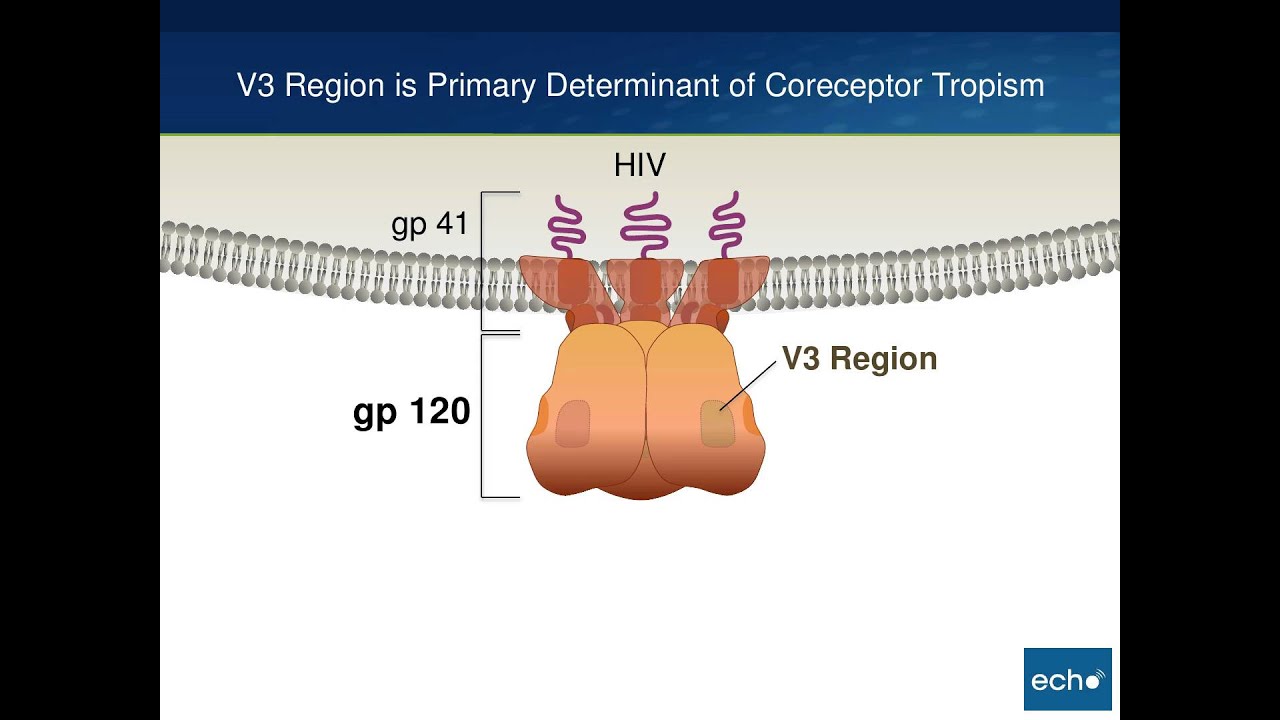
On the host cell surface, the primary surface receptors involved during entry are the CD4 receptor and the chemokine coreceptors, either CCR5 or CXCR4. The key HIV component in cell entry is the envelope glycoprotein, which consists of the outer gp120 subunit and the inner folded gp41 subunit. The gp120 subunit has two regions that are particularly important in entry: the V3 region and an outer groove known as the CD4 binding site.
The initial contact between HIV and the host cell typically involves binding of the HIV gp120 with the host cell CD4 receptor, and this occurs at the CD4 binding site. This initial binding is considered the first phase of HIV entry and is referred to as attachment. This attachment stimulates the HIV envelope V3 region to migrate fluidly to the distal region of gp120, stimulating binding between the HIV envelope region and the host chemokine coreceptor, most often, the CCR5 coreceptor.
This interaction is referred to as coreceptor binding and is considered the second phase in HIV entry. The coreceptor binding triggers the folded gp41 to spring open and latch onto the surface of the host cell. As this occurs, the gp120 subunits are cast off and they're no longer involved in viral entry.
The gp41 subunit has a unique ability to fold back on itself with the two main gp41 domains sticking together like Velcro. This folding triggers the attached virion to roll across the surface of the host cell membrane, creating more interactions between the HIV envelope and the host receptors. As this occurs, the multiple gp41 folding events pulls the HIV membrane and the host membranes together, initiating the membrane fusion process.
As this process continues, the HIV membrane and the HIV matrix shell fully merge with the host cell membrane. This fusion of the membranes is quickly followed by the migration of the HIV core into the host cell. The HIV core is also referred to as the HIV capsid or HIV capsid core.
As this process continues, the HIV core moves further into the host cell. And with the full merger of HIV and host membranes, the HIV core has moved entirely inside of the host cell. At this point, the third and final phase of HIV entry fusion is complete.
Let s take a look at the next step in the HIV life cycle: the cytoplasmic transport, and nuclear import of the HIV core. Until recently, it was thought the HIV core dissolved soon after it enters the host cell cytoplasm. Current research suggests the HIV core usually remains intact, or mostly intact, until after entering the host cell nucleus.
This view is a cross-section of the HIV core and note several of the key components inside the core that need to be delivered into the nucleus, most importantly the genomic HIV RNA and the HIV enzymes integrase and reverse transcriptase. The cytoplasmic transport of the HIV core to the nucleus utilizes the host microtubule system. The transport via the microtubules is a dynamic process that involves multiple host proteins including dynein and kinesin.
Dynein accelerates transport toward the nucleus, whereas kinesin promotes movement away from the nucleus. During cytoplasmic transport, the intact cone-shaped HIV core moves along the microtubule until it reaches the outer region of the host cell nucleus. For the core to make its way into the nucleus, it must pass through the nuclear pore.
The nuclear pore is a complex structure that consists of outer cytoplasmic ring, a central channel, a nuclear ring, and a nuclear basket. In the initial step of nuclear import, the HIV capsid proteins on the more narrow end of the HIV core interact with the nuclear pore and nuclear protein 358, resulting in docking of the HIV core on the cytoplasmic side of the nuclear pore complex. As the HIV core penetrates into the central channel of the nuclear pore, it encounters several additional host cell nuclear transport proteins, as shown in orange and yellow here.
These proteins assist the HIV core in moving through the nuclear pore. As the HIV core begins to merge into the nucleus, it binds with the host protein CPSF6, which stands for cleavage and polyadenylation specificity factor 6. This protein also assists the HIV core in movement through the nuclear pore.
As the HIV core advances further through the pore, it becomes more heavily coated with the CPSF6 protein. At this stage, the cone-shaped HIV core is starting to lose some of its structural integrity. As the core exits a nuclear pore and fully enters into the nucleus, there's major structural breakdown of the HIV core.
Once completely inside the nucleus, the HIV core undergoes further disassembly, releasing the HIV DNA and HIV integrase enzyme, thus completing the cytoplasmic transport and nuclear import step. Now let's go back and look at HIV reverse transcription. This process is occurring during the transport of the HIV core to the nucleus.
Reverse transcription is the step involving the conversion of HIV RNA into HIV DNA. Soon after the HIV core enters the host cell the reverse transcription of genomic HIV RNA begins, and it starts inside of the HIV core. The key concept in HIV reverse transcription is that the genomic HIV RNA is converted into HIV DNA.
The enzyme that catalyzes this process is reverse transcriptase. The reverse transcriptase protein has two key enzymatic regions: the polymerase region, which facilitates the building out of the new DNA strand and ribonuclease H, which degrades some of the unwanted intermediate RNA and DNA segments during reverse transcription. The building blocks used to generate the HIV DNA are taken from the intracellular pool of human nucleotides.
Each nucleotide consists of a nitrogenous base, a pentose sugar, and a phosphate. Nucleotides adenine, guanine, and cytosine can be used to make either DNA or RNA. Thymidine is only used for DNA synthesis and uracil only for RNA synthesis.
For conceptual purposes in the following illustrations, I will be using a simplified representation of the human nucleotides, as shown in the bottom row here. Although the process of reverse transcription is extremely complicated, the basic idea is that reverse transcriptase orchestrates the binding of the host cell nucleotides to the complementary nucleotides on the HIV RNA template strand. This process continues with the addition of one nucleotide at a time, eventually generating double-stranded HIV DNA.
For this process, the original HIV RNA strand is referred to as the template strand and the newly forming strand as the primer strand. Now let's step back and look at where reverse transcription is taking place. As noted earlier, most of the reverse transcription takes place inside of the HIV core, starting soon after the core enters the cytoplasm of the host cell.
As the HIV core migrates toward the nucleus, early HIV reverse transcription is underway, generating intermediate hybrids consisting of HIV RNA and single-stranded HIV DNA. As the HIV core reaches the nuclear pore, the reverse transcription process is further along, and more single-stranded HIV DNA is likely forming. The HIV core enters into the host nucleus with the core entirely, or mostly, intact.
At this point, HIV double-stranded DNA has started to form. The double-stranded DNA is structurally less flexible than HIV RNA, and this rigid DNA is believed to cause some disruption in the outer shell of the HIV core. As more of the stiffer double-stranded HIV DNA continues to form, the structural Integrity of the HIV core breaks up further.
Eventually, the HIV core breaks down completely, releasing the HIV DNA into the host cell nucleus. The HIV DNA is now ready for integration, the next phase in the HIV replication cycle. Okay, now let's look at HIV integration, the process whereby the newly formed HIV DNA is stitched into the human DNA.
The HIV integration process begins soon after the HIV DNA synthesis has been completed. For this step to get underway, the HIV integrase enzyme must bind to the ends of the newly formed HIV DNA, as shown here. The complex of the HIV integrase enzyme bound to the HIV DNA is referred to as the intasome.
Most models suggest integrase initially binds to the ends of the viral DNA as separate dimers but at some point prior to integration The dimers join to form a tetramer intasome structure. In the illustrations that follow, for conceptual purposes, I will primarily show the dimer into some confirmation. The first phase of HIV integration is referred to as 3-prime processing.
This involves the integrase enzyme cleaving two nucleotides off the 3-prime ends of the HIV DNA. This process generates reactive 3-prime hydroxyl groups on the ends of the HIV DNA, which prepares it for insertion into the host DNA. The second phase in HIV integration is referred to as strand transfer.
During this phase, the HIV integration enzyme catalyzes the insertion and transfer of the strand of HIV DNA into the host cell DNA. This initial attack, and the strand transfer reaction occurs after the HIV intasome docks onto the host cell DNA. The active component of the integrase enzyme is the catalytic core region.
This close-up view shows the multi-pronged attack on the host DNA by the reactive 3-prime hydroxyl ends of the HIV DNA in tandem with a catalytic core region of the HIV integrase enzyme. A second attack by HIV takes place on the opposite strand of the host DNA, typically 4-6 base pairs apart from the initial side of attack. These two sequential reactions cause separation and the strands of the host DNA.
The host DNA strands separate further and the HIV integrase enzymes are released as they are no longer needed. The newly formed HIV host DNA hybrid straightens out but unpaired regions of DNA remain at the host HIV DNA junctions. These regions are referred to as DNA gaps.
The host cell enzymes step in and repair these gaps using host nucleotides; additional host enzymes smooth over the newly formed junctions as part of this gap repair process. At this point, the transfer and integration of the HIV DNA into the host DNA is considered complete, and the fully integrated HIV DNA is referred to as the HIV provirus or proviral DNA. Now, let's take a look at HIV transcription, which should not be confused with HIV reverse transcription.
The HIV transcription step involves converting the HIV proviral DNA into HIV RNA. Similar to other steps in the HIV life cycle, HIV takes advantage of the existing elements in the human cell, essentially hijacking the host system that normally transcribes human DNA into human RNA. HIV transcription can begin only after the HIV DNA has been successfully integrated into the host DNA.
The host cell RNA polymerase is the key enzyme involved in transcribing the HIV DNA into HIV RNA. Similar to HIV reverse transcription, the HIV transcription utilizes the host nucleotide pool but, in this case, the nucleotides are used to generate RNA, not DNA. HIV transcription produces two types of HIV RNA: genomic RNA, which is used as the genome for a new HIV particle and messenger RNA, or mRNA for short, which is used to synthesize HIV proteins.
The active phase of transcription begins after the host cell polymerase binds to the promoter region of the HIV DNA. The HIV transcription begins, but soon thereafter, it pauses. But binding of the HIV accessory protein transactivator of transcription, or tat for short, overrides this pause, and the efficiency of the viral transcription dramatically increases, particularly with the full length genomic HIV RNA.
The HIV that is generated can also be grouped into three categories based on the splicing that occurs: 1) full-length RNA, 2) partially-spliced mRNA, and 3) fully-spliced mRNA. The newly formed RNA leaves the host cell nucleus. The the unspliced genomic HIV RNA migrates to the cell's surface where it is packaged into a new virion.
The mRNA transcripts that migrate into the cytoplasm serve as the templates for the synthesis of HIV proteins. At this point, the next stage in the HIV life cycle, translation, can begin. The HIV translation process occurs in the cytoplasm, and this is the step in the life cycle where HIV mRNA is translated into 15 different HIV proteins.
During HIV translation, an extensive array of the host cell translation machinery is utilized, including ribosomes, transfer RNAs, amino acids, and additional factors required for initiation, elongation, and termination of protein synthesis. The fully-spliced HIV mRNA is often first to undergo translation, generating the HIV regulatory proteins rev and tat along with the HIV accessory protein nef. Much of the rev that is synthesized rapidly migrates back into the host cell nucleus where it enhances the export of partially-spliced and unspliced HIV mRNA into the cytoplasm.
Similarly HIV tat is also shoveled back into the nucleus where, as discussed earlier, tat binds to the HIV DNA and increases the efficiency of HIV transcription. The partially-spliced HIV mRNA transcripts are used to synthesize the HIV accessory proteins vif, vpr, and vpu as well as the HIV envelope precursor protein gp160. Let's take a closer look at the synthesis and processing of the HIV envelope protein along with vpu.
This process occurs along the host cell endoplasmic reticulum with the newly forming protein secreted into the lumen of the endoplasmic reticulum. Once translation of the vpu envelope mRNA is complete, the accessory protein vpu is released, but the HIV envelope precursor protein gp160 remains attached to the endoplasmic reticulum membrane via the signal peptide at its N-terminal end. During the translation of the envelope precursor protein, the outer component is coded with multiple oligosaccharide side chains, which, in essence, generates a glycan shield around the envelope.
For ease of understanding, the following images will not show this glycan shield. In the next phase of processing the HIV envelope precursor protein, the host enzyme signal peptidase cleaves the HIV signal peptide, which frees the N-terminal end of the envelope protein from the endoplasmic reticulum membrane: this protein is now referred to as an HIV gp160 monomer. The gp160 monomers that are generated unite along the endoplasmic reticulum membrane, mostly as trimers but some as dimers.
The formation of these complex triggers trafficking of the gp160 to the golgi complex. Once inside the golgi, the host enzyme furin cleaves each of the gp160 proteins into separate gp120 and gp41 subunits. After the furin cleavage, the free gp41 subunits transform into a folded conformation and then closely associate with the gp120 subunits through a noncovalent bond.
The newly formed gp120-gp41 pairs form a mature trimer, which is also known as the envelope glycoprotein spike. Note that each trimer consists of gp120-gp41 pairs, as shown schematically here in the inset, with each gp120 bound to gp41 by a noncovalent bond. The last mRNAs to review in the transcription step are the full-length unspliced mRNA transcripts, which are translated into large gag and gag-pol polyproteins.
Conceptually, it is important to understand how both of these large polyproteins are synthesized from the same full-length unspliced mRNA. About 95% of the time only part of this unspliced mRNA is translated, which generates the HIV gag polyprotein but about 5% of the time, after most of the way through translating the gag polyprotein, the ribosome slips back one nucleotide. This -1 ribosomal frame shift resets the reading frame and this extends the mRNA translation, generating the very large gag-pol polyprotein.
Overall, about 20 gag polyproteins are formed for every one gag-pol polyprotein. The next to last phase in the HIV lifecycle is HIV assembly. During HIV assembly, the structural and functional components of HIV migrate to the surface of the host cell and begin to assemble at the cell membrane.
Now let's take a look at the assembly of three HIV components: genomic RNA, envelope proteins, and the large gag and gag polyproteins. For each new virion that is formed, two strands of genomic HIV RNA are needed. After the genomic HIV RNA migrates to the cell surface, it becomes intertwined with the HIV gag polyproteins.
For the next component of assembly, let's look at the HIV envelope. As noted earlier, the final processing of the HIV envelope protein occurs in the golgi. The HIV envelope trimers are then shuttled from the golgi to the cell surface by cellular vesicles.
After reaching the cell surface, the envelope protein undergoes two fates: some of the envelope proteins will become part of the newly assembling HIV particle, and some will undergo recycling, either returning to the cell surface or undergoing degradation within lysosomes. Last, but not least, an assembly process. The HIV polyproteins gag and gag-pol play a critical role in the structure and function of the newly forming virion.
These large polyproteins migrate toward the host cell membrane in clusters with their most-N-terminal component, the matrix protein, facing toward the cell membrane. As the gag and gag-pol polyproteins reach the cell's surface, the matrix proteins bind to the inner surface of the host membrane. Let's take a closer look at the matrix in the host membrane interaction.
This interaction is initiated when a highly basic region of the matrix protein, shown in purple here, associates with the specific host membrane lipid known as phosphatidylinositol-4,5-bisphosphate. This interaction stimulates the matrix myristyl group to transform from its baseline, sequestered conformation, as shown here, to an exposed conformation that inserts into the host lipid bilayer, firmly interlocking the matrix proteins with the host cell membrane. The change in the matrix myristyl conformation is also associated with the flipping of a host membrane lipid chain into the matrix protein, which further strengthens the binding between the matrix protein and the host membrane.
The interlocking of the matrix protein with the inner surface of the membrane also provides stabilization of the envelope protein binding to the host cell membrane. And last for this stage, the HIV accessory protein vpr binds to several gag proteins at the p6 terminal end, essentially docking on to get a ride into the newly forming virion. The final step in the HIV life cycle is HIV budding and maturation.
As the assembly process continues, the number of gag polyproteins increases exponentially at the host membrane surface until a plateau is reached, triggering the host membrane to pinch inward, as shown on the left here. During this budding process, the gag proteins align radially like spokes in a wheel with the matrix bound to the membrane. The budding process is facilitated by the most-distal gag protein p6.
One region of p6 near the N-terminal region is critical in this process: a 4-amino acid sequence located at residues 7 through 10. This region is referred to as the PTAP sequence, and these four amino acids interact with and activate the endosomal sorting complex that is required for transport; this is abbreviated as ESCRT. The ESCRT complexes, which are not shown here, are a group of host proteins that facilitate viral budding.
As the immature virion fully separates from the host cell, it is important to note that the lipid membrane and the newly forming HIV particle have been acquired from the host membrane. At this point, the HIV maturation phase begins with the clustering of the gag polyproteins. The clustering is largely driven by lateral binding between the capsid proteins with other capsid proteins.
This close-up view shows the clustering of multiple gag polyproteins. At this early phase in maturation, the two strands of genomic HIV RNA are located toward the center of the immature virion and are tightly aligned with the nucleocapsid protein. More extensive capsid-capsid binding leads to the formation of capsid hexamers, as shown in the inset cross-sectional view here.
Occasional capsid pentamers are also formed. Along with the capsid clustering, the matrix proteins group as trimers. At this stage, the HIV protease enzyme is autocleaved from the gag-pol polyprotein.
The newly released HIV protease enzyme begins processing the gag and the gag-pol polyproteins. The protease enzyme cleaves the polyproteins in a predictable and consistent order, as shown here with the sequential five-step processing of the gag polyprotein. As the protein processing occurs, the HIV matrix proteins remain bound to the HIV membrane but the other cleaved proteins become untethered and begin to form the internal structure of the virion.
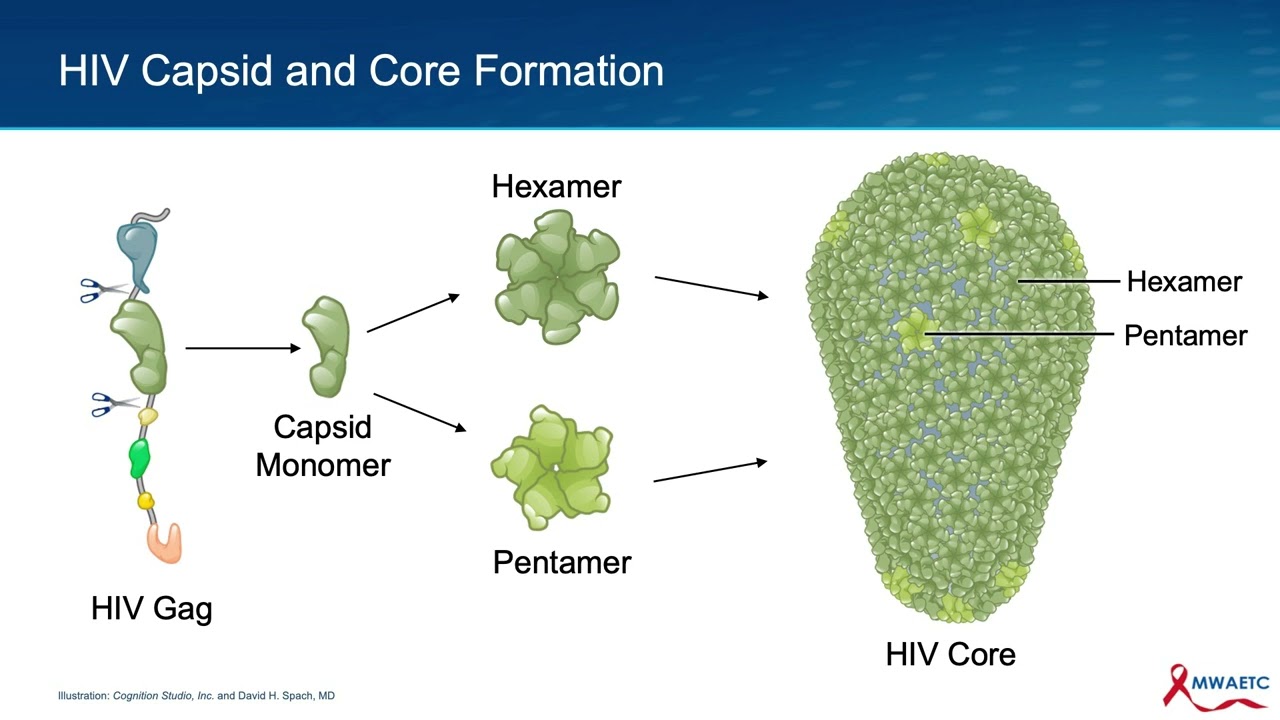
The key structural event and the formation of the HIV core is the binding of the HIV capsid protein clusters to each other. As this core begins to form, the two strands of HIV RNA and several key enzymes are positioned inside of this newly forming structure. The process of core formation is complex and begins with capsid monomer proteins grouping into hexamers and pentamer units.
As noted earlier, these polymers initially form as part of the gag and gag-pol polyproteins. But after protease freeze the capsid proteins from other elements of gag and gag-pol polyproteins, the capsid polymers can further group into a lattice-like structure, the early HIV core. The process continues with the complete assembly of the hexamer and pentamer units to eventually form the cone-shaped HIV core.
The fully formed HIV core structure is comprised of about 250 hexamers, shown in darker green, and 12 pentamers, shown in lighter green. The generation of the fully formed core marks the completion of the viral replication process. The final product is a mature virion that is capable of infecting new cells.
Hopefully, as a result of this talk you have an appreciation for how HIV enters a host cell and takes over the inner workings of the cell to generate new HIV particles, gradually turning the cell into a virus-making factory, eventually depleting the host cell of many of its essential elements, which results in the demise of the cell. In closing, during this talk I've examined in detail the essential steps required by HIV to produce new virions. I hope this talk has provided you with a deeper understanding of the HIV life cycle.
Thank You. The production of this National HIV Curriculum Mini-Lecture was supported by funding from the Health Resources and Services Administration.